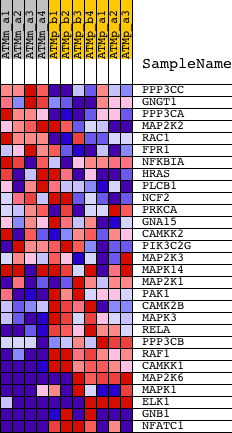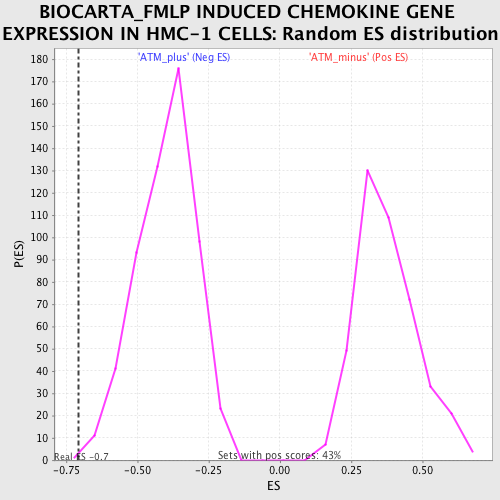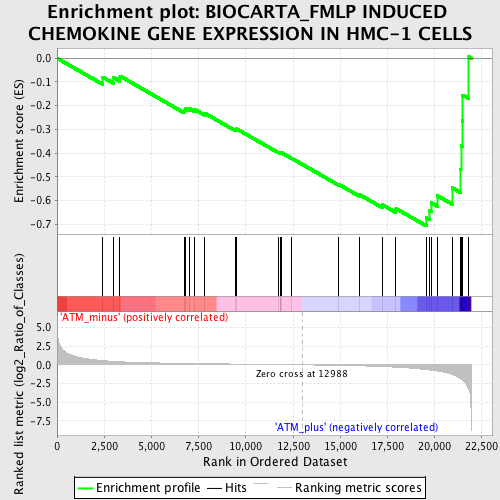
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | BIOCARTA_FMLP INDUCED CHEMOKINE GENE EXPRESSION IN HMC-1 CELLS |
| Enrichment Score (ES) | -0.7069288 |
| Normalized Enrichment Score (NES) | -1.7755346 |
| Nominal p-value | 0.0017391305 |
| FDR q-value | 0.07349958 |
| FWER p-Value | 0.388 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PPP3CC | 1420743_a_at 1430025_at | 2406 | 0.566 | -0.0794 | No | ||
| 2 | GNGT1 | 1425167_a_at 1425168_at 1425219_x_at 1451633_a_at | 2967 | 0.454 | -0.0805 | No | ||
| 3 | PPP3CA | 1426401_at 1438478_a_at 1440051_at 1452056_s_at 1458495_at | 3329 | 0.407 | -0.0751 | No | ||
| 4 | MAP2K2 | 1415974_at 1443436_at 1460636_at | 6734 | 0.192 | -0.2202 | No | ||
| 5 | RAC1 | 1423734_at 1437674_at 1451086_s_at | 6772 | 0.190 | -0.2116 | No | ||
| 6 | FPR1 | 1450808_at | 7003 | 0.182 | -0.2123 | No | ||
| 7 | NFKBIA | 1420088_at 1420089_at 1438157_s_at 1448306_at 1449731_s_at | 7278 | 0.173 | -0.2155 | No | ||
| 8 | HRAS | 1422407_s_at 1424132_at | 7807 | 0.155 | -0.2312 | No | ||
| 9 | PLCB1 | 1421170_a_at 1425600_a_at 1425781_a_at 1425782_at 1435043_at 1447004_at | 9454 | 0.104 | -0.3008 | No | ||
| 10 | NCF2 | 1448561_at | 9514 | 0.102 | -0.2980 | No | ||
| 11 | PRKCA | 1427562_a_at 1443069_at 1445028_at 1445395_at 1446598_at 1450945_at | 11704 | 0.039 | -0.3958 | No | ||
| 12 | GNA15 | 1421302_a_at | 11821 | 0.036 | -0.3992 | No | ||
| 13 | CAMKK2 | 1424474_a_at 1424475_at 1424476_at 1455401_at | 11833 | 0.035 | -0.3978 | No | ||
| 14 | PIK3C2G | 1421704_a_at | 11868 | 0.034 | -0.3975 | No | ||
| 15 | MAP2K3 | 1425456_a_at 1451714_a_at | 12398 | 0.018 | -0.4206 | No | ||
| 16 | MAPK14 | 1416703_at 1416704_at 1426104_at 1442364_at 1451927_a_at 1459617_at | 14906 | -0.071 | -0.5312 | No | ||
| 17 | MAP2K1 | 1416351_at | 16004 | -0.128 | -0.5744 | No | ||
| 18 | PAK1 | 1420979_at 1420980_at 1450070_s_at 1459029_at | 17237 | -0.231 | -0.6183 | No | ||
| 19 | CAMK2B | 1448676_at 1455869_at | 17945 | -0.319 | -0.6334 | No | ||
| 20 | MAPK3 | 1427060_at | 19557 | -0.635 | -0.6728 | Yes | ||
| 21 | RELA | 1419536_a_at | 19723 | -0.683 | -0.6435 | Yes | ||
| 22 | PPP3CB | 1427468_at 1428473_at 1428474_at 1433835_at 1446149_at 1459378_at | 19815 | -0.712 | -0.6093 | Yes | ||
| 23 | RAF1 | 1416078_s_at 1425419_a_at | 20130 | -0.828 | -0.5791 | Yes | ||
| 24 | CAMKK1 | 1418954_at | 20930 | -1.287 | -0.5463 | Yes | ||
| 25 | MAP2K6 | 1426850_a_at 1459292_at | 21369 | -1.830 | -0.4678 | Yes | ||
| 26 | MAPK1 | 1419568_at 1426585_s_at 1442876_at 1453104_at | 21390 | -1.864 | -0.3684 | Yes | ||
| 27 | ELK1 | 1421896_at 1421897_at 1446390_at | 21455 | -1.991 | -0.2641 | Yes | ||
| 28 | GNB1 | 1417432_a_at 1425908_at 1454696_at 1459442_at | 21482 | -2.043 | -0.1553 | Yes | ||
| 29 | NFATC1 | 1417621_at 1425761_a_at 1428479_at 1447084_at 1447085_s_at | 21810 | -3.267 | 0.0057 | Yes |